missing translation for 'onlineSavingsMsg'
Learn More
Learn More
Invitrogen™ JAKMIP2 Polyclonal Antibody


Description
Immunogen sequence: RETEKQCKPLL ERNKCLAKRN DELMVSLQRM EEKLKAVTKE NSEMREKITS HPPLKKLKSL NDLDQANEEQ Highest antigen sequence identity to the following orthologs - mouse 100%, rat 99%.
JAKMIP2 (janus kinase and microtubule-interacting protein 2), also known as NECC1 (neuroendocrine long coiled-coil protein 1), CTCL tumor antigen HD-CL-04, JAMIP2 or KIAA0555, is a 810 amino acid protein belonging to the JAKMIP family. Localizing to the Golgi apparatus, JAKMIP2 is high expressed in brain, with moderate levels of expression found in thymus, spleen and lung. Existing as three alternatively spliced isoforms, the gene encoding JAKMIP2 maps to human chromosome 5q32 and mouse chromosome 18 B3. Chromosome 5 contains 181 million base pairs and comprises nearly 6% of the human genome. Deletion of the p arm of chromosome 5 leads to Cri du chat syndrome, while deletion of the q arm or of chromosome 5 altogether is common in therapy-related acute myelogenous leukemias and myelodysplastic syndrome.

Specifications
Specifications
| Antigen | JAKMIP2 |
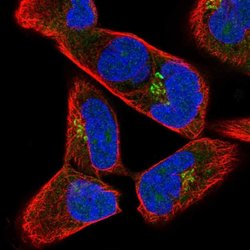
| Applications | Immunocytochemistry |
| Classification | Polyclonal |
| Concentration | 0.2 mg/mL |
| Conjugate | Unconjugated |
| Formulation | PBS with 40% glycerol and 0.02% sodium azide; pH 7.2 |
| Gene | Jakmip2 |
| Gene Accession No. | Q96AA8 |
| Gene Alias | 6430702L21Rik; AI850334; CTCL tumor antigen HD-CL-04; D930046L20Rik; Jak and microtubule interacting protein 2; JAKMIP2; JAMIP2; janus kinase and microtubule interacting protein 2; janus kinase and microtubule-interacting protein 2; KIAA0555; NECC1; Neuroendocrine long coiled-coil protein 1; RGD1559742 |
| Gene Symbols | Jakmip2 |
| Show More |
Product Title
By clicking Submit, you acknowledge that you may be contacted by Fisher Scientific in regards to the feedback you have provided in this form. We will not share your information for any other purposes. All contact information provided shall also be maintained in accordance with our Privacy Policy.
Spot an opportunity for improvement?